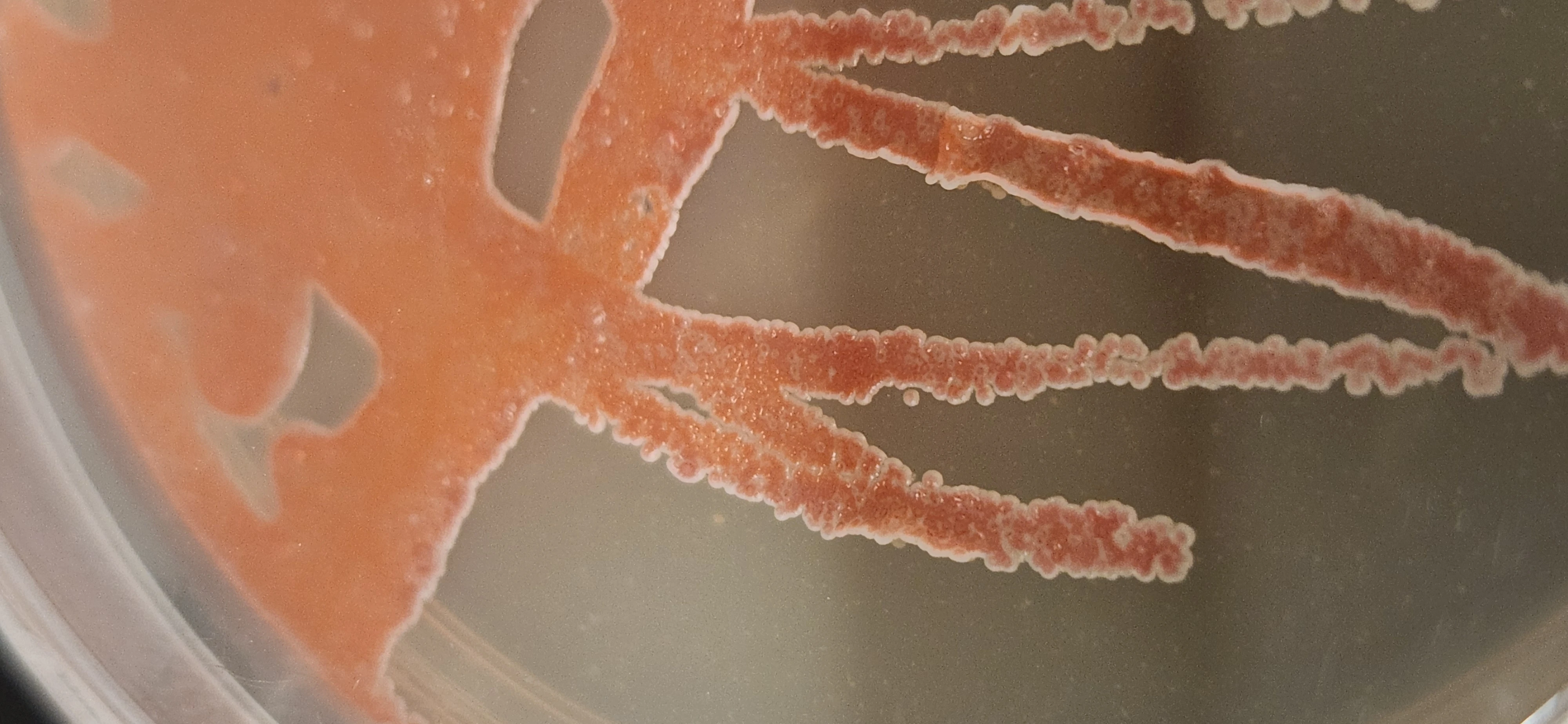
The World's Largest BGC Atlas
We've efficiently turned tens of thousands of compute hours into 5.2 million BGCs (27.8M unique proteins) from 750,000 genomes and metagenomes. These have been clustered with proprietary software that operates at orders of magnitude greater speed and scale than currently reported tools.
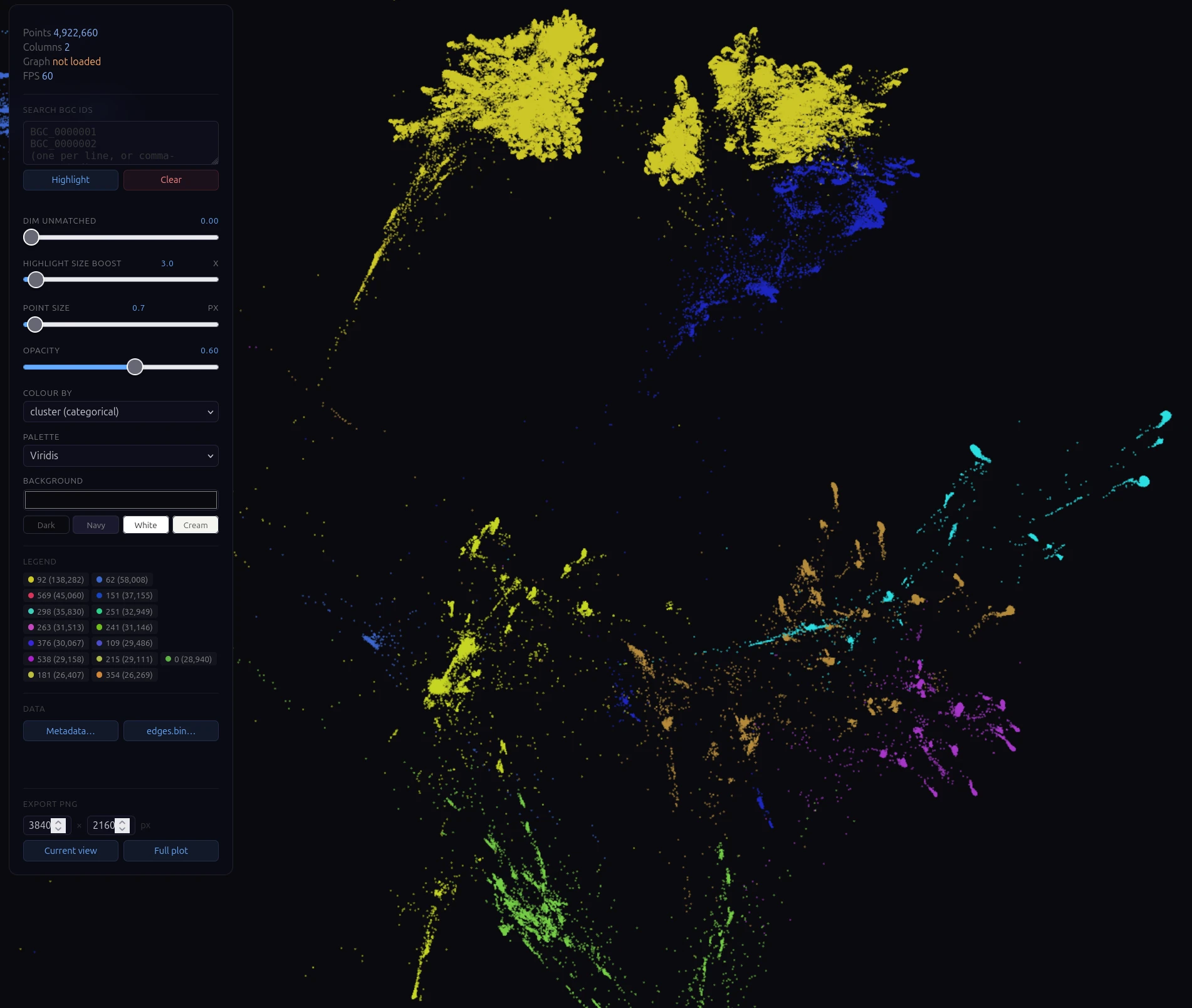
When BioPredictor surfaces a BGC of interest, we instantly compare it to all 5.2 million BGCs in the atlas, contextualizing novelty, chemical class, and taxonomic distribution in a single view, and flagging anything that maps onto a previously characterized cluster.
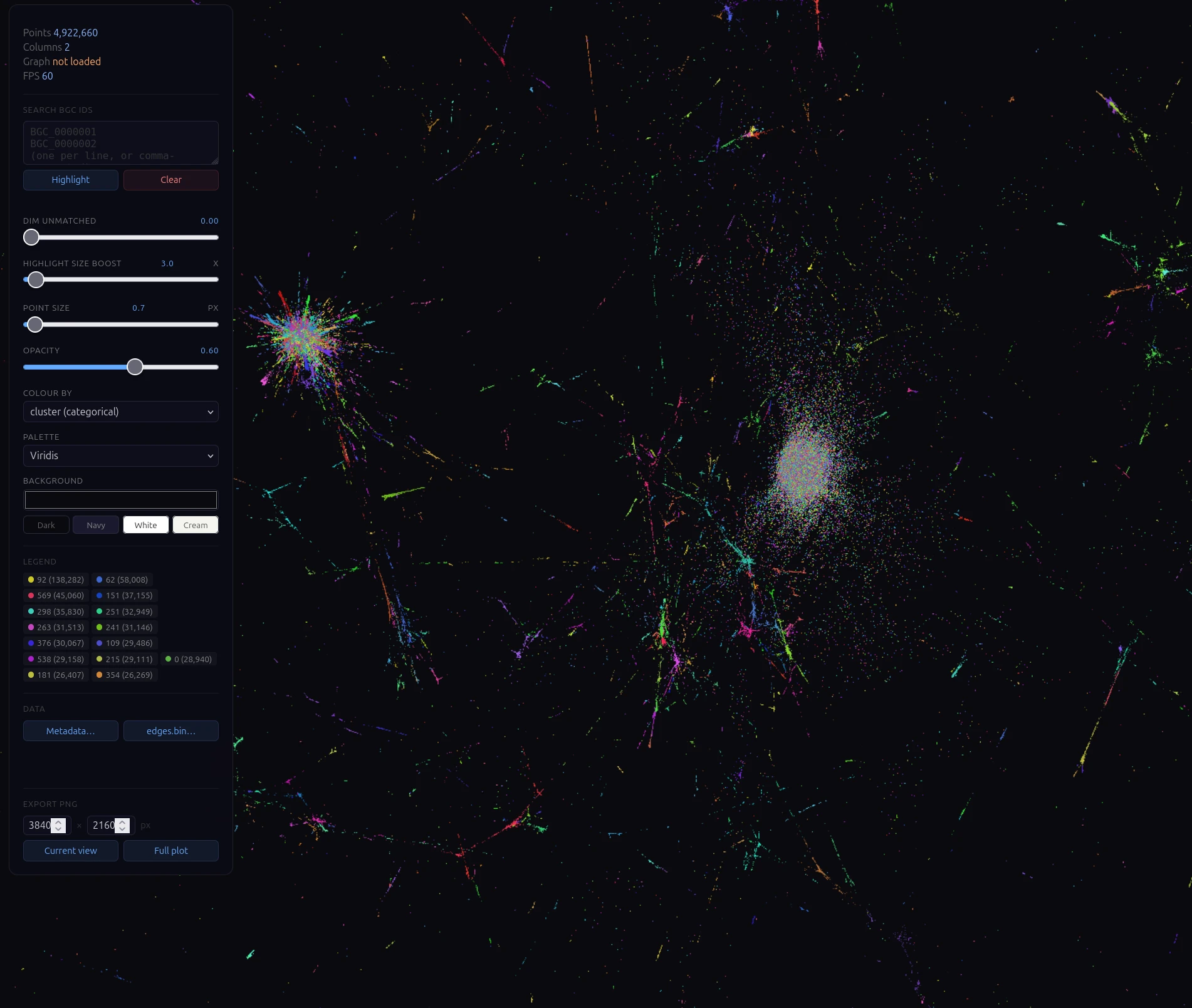
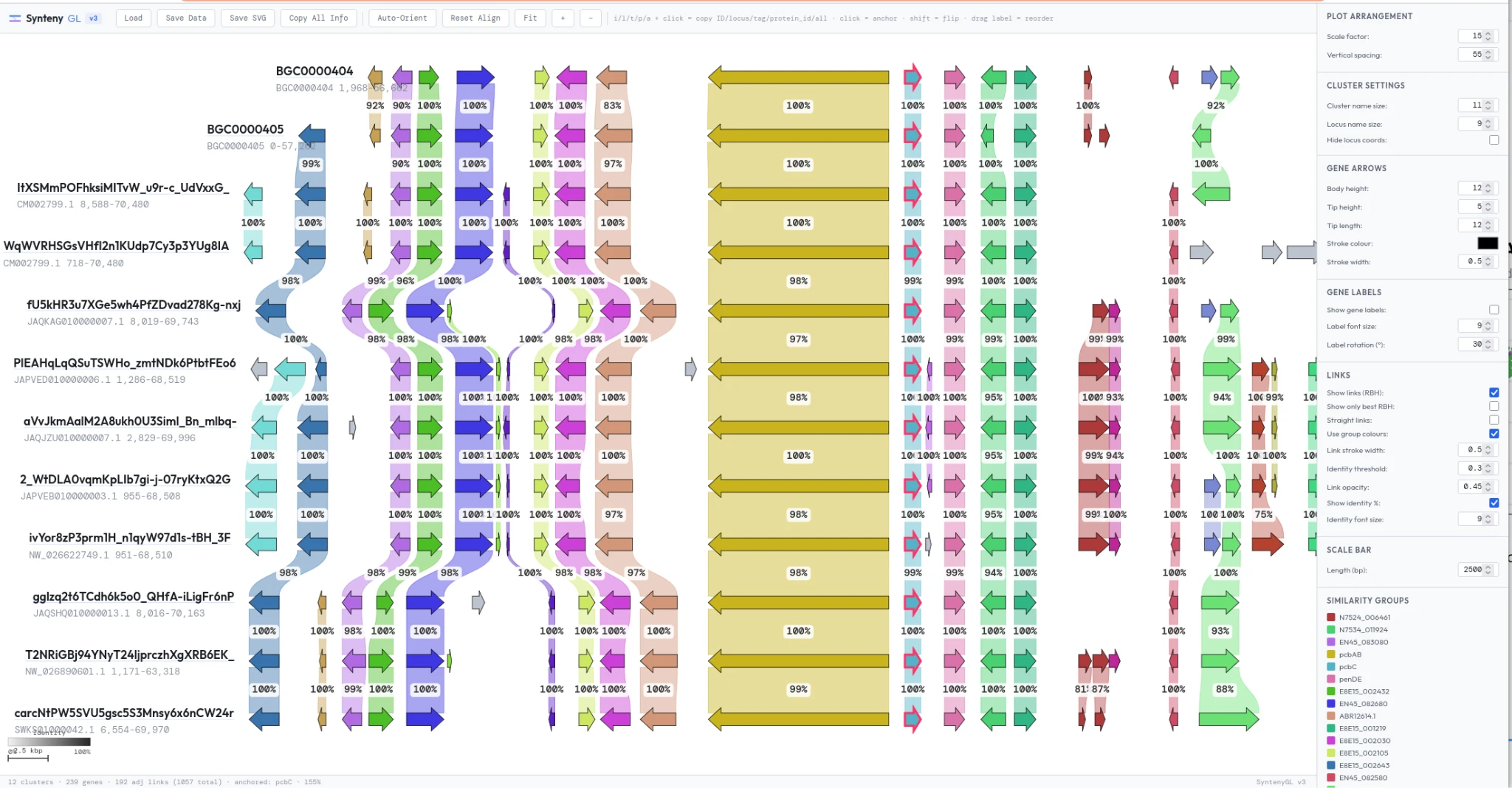
Selecting nodes in the atlas passes BGCs to our synteny analysis workflow. In seconds, BGCs are pulled from the cloud and rendered in a single interactive visualization, revealing shared and unique genes, domain architectures, and overall organization at a glance.